Abstract
Agmial is an integrated system for bacterial genome annotation. It has been developed with the following requirements in mind:
- maximize annotation automation,
- ability to work on draft sequences,
- keep the program modular and extensible,
- choose informatics and bioinformatics standards when possible (e.g. Web Services, RDBMS, ...)
- distribute the program under an open-source license.
Demonstration
A demonstration version of agmial is available here
(artemis).
The login is agmialRead and the password
agmialRead
A version of agmial with some public genomes re-annoted is available here
(artemis).
References
- K. Bryson, V. Loux, R. Bossy, P. Nicolas, S. Chaillou, M. van de Guchte, S. Penaud, E. Maguin JF. Gibrat. AGMIAL: implementing an annotation strategy for prokaryote genomes as a distributed system. Nucleic Acids Research. Jul 2006. [Pubmed]
-
Eric Duchaud, Mekki Boussaha, Valentin Loux, Jean-François Bernardet, Christian Michel, Brigitte Kerouault, Stanislas Mondot, Pierre Nicolas, Robert Bossy, Christophe Caron, Philippe Bessieres, Jean-François Gibrat, Stéphane Claverol, Fabien Dumetz, Michel Le Hénaff, Abdenour Benmansour. Complete genome sequence of the fish pathogen Flavobacterium psychrophilumNat Biotechnol2007 [Pubmed]
- M. van de Guchte , S. Penaud , C. Grimaldi , V. Barbe , K. Bryson, P. Nicolas, C. Robert, S. Oztas, S. Mangenot, A. Couloux, V. Loux<, R. Dervyn, R. Bossy, A. Bolotin, JM. Batto, T. Walunas, JF. Gibrat, P. Bessieres, J. Weissenbach, SD. Ehrlich, E. Maguin.The complete genome sequence of Lactobacillus bulgaricus reveals extensive and ongoing reductive evolution. Proceedings of the National Academy of Sciences of the United States of America. Jun 2006. [Pubmed]
- S. Chaillou, M. Champomier-Verges, M. Cornet, A-M Crutz Le Coq, A-M.Dudez, V. Martin, S. Beaufils, E. Darbon-Rongere, R. Bossy, V. Loux and M. Zagorec. The complete genome sequence of the meat-borne lactic acid bacterium Lactobacillus sakei 23K. Nature Biotechnology December 2005. [Pubmed]
Collaborations
Agmial is used or has been used to annotate :
- Flavobacterium psychrophilum and other flavobacter (Virologie et Immunologie Molüķculaire,INRA Jouy en Josas , SA Department ) [Public server] [Private server] [Artemis].
- Lactobacillus bulgaricus (Genetique Microbienne,INRA Jouy en Josas , MICA department)[Private server] [Artemis].
- Lactobacillus sakei(Flore Lactique et Environnement Carnüķ ,INRA Jouy en Josas , MICA department) [Public server] [Private server] [Artemis].
- Propionibacterium freudenrichii( Science et technologie du lait et de l'oeuf ,INRA Rennes , MICA department) [Private server] [Artemis].
- Staphylococcus xylosus( Microbiologie MIC,INRA Theix , MICA department) [Private server] [ [Artemis].
- Streptococcus salivarius JIM8777 and JIM8780 (Genetique Microbienne,INRA Jouy en Josas , MICA department)[Private server] [Artemis].
- Arthrobacter arilaitensisand other genomes( GMPA,INRA Grignon , MICA department) [Private server] [ [Artemis].
- And other private genomes ...
Trouble with Artemis? Check Artemis User Guide and Faq
Contacts
Jean-François Gibrat ,Valentin Loux.
Architecture
The Agmial architecture involves two independent modules: the Contig Analysis Module (CAM) and the Protein Analysis Module (PAM). They are respectively responsible for storing and applying bioinformatics tools on DNA and proteins sequences. The system currently integrates 20 different third-party programs including the SHOW gene detector, Blast and the Interpro suite. Thanks to the modular architecture of Agmial, the addition of new tools represents a marginal effort of development.
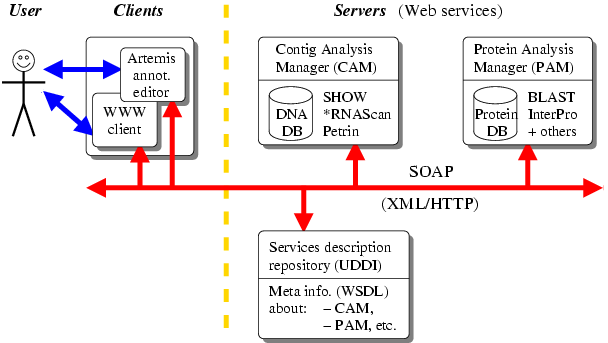
In order to synchronize the annotations of proteins on PAM and their corresponding CDS on CAM, modules communicate through the SOAP protocol; as a matter of fact both modules are web service providers. The end-user may interact with the system through either a web browser, either a Web Services client: we currently use a modified version of Artemis.
Usage data flow
The close cooperation between the system and the experts leads to an accurate annotation of the genome. The end-user is usually a team of annotators who works on a specific organism: they input the draft genome sequence, CAM searches for CDSs, passes the translated proteins to PAM which looks for homologies and domains.
After all the processing, the system suggests a functional annotation for each protein, then the annotators validate and refine it. The following diagram represents the information exchanged during the annotation process:
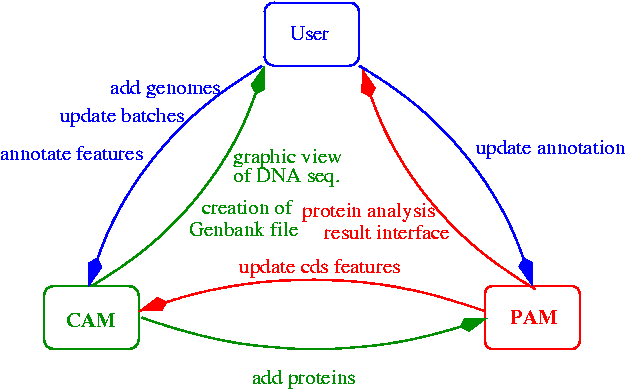
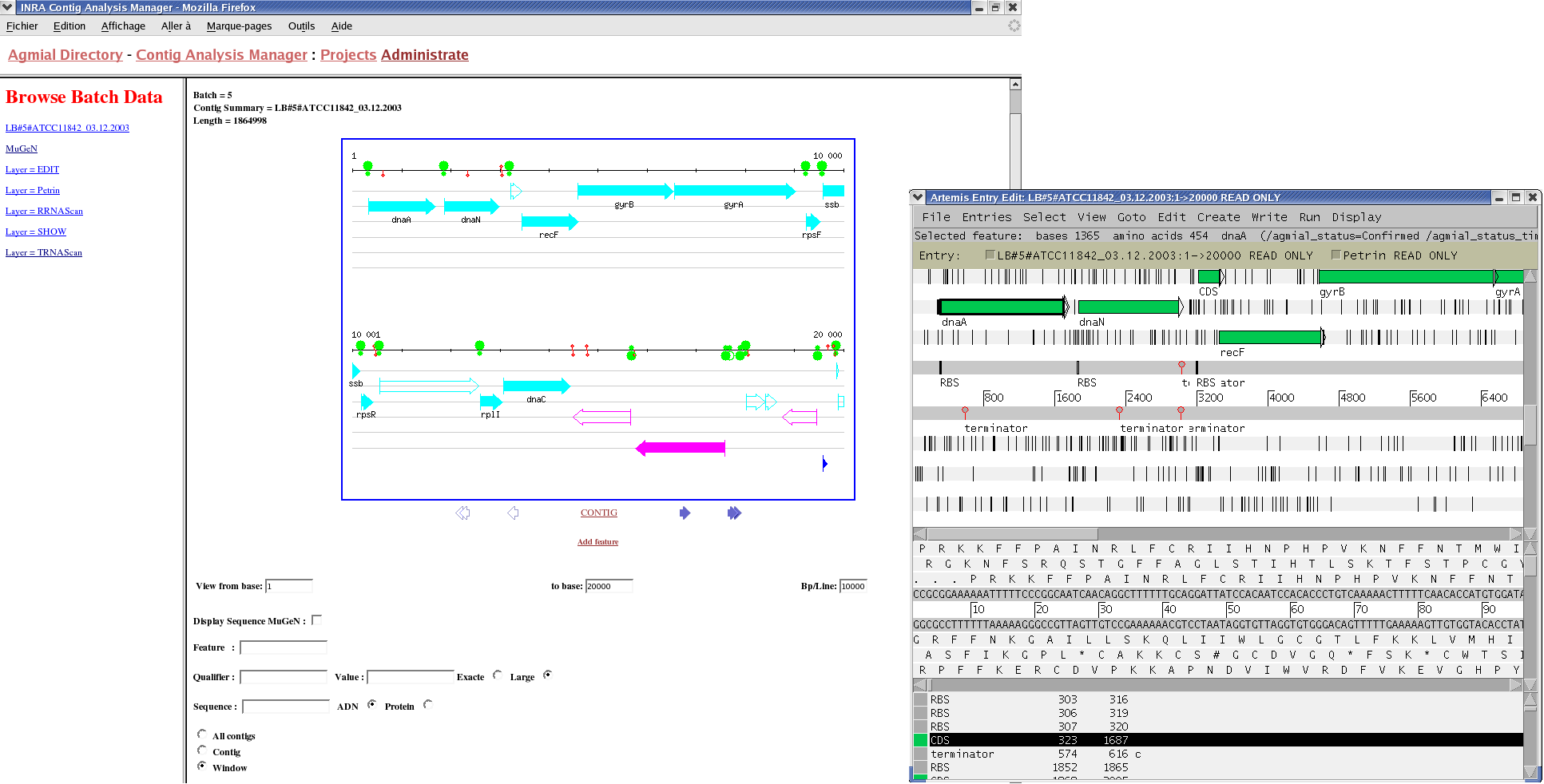
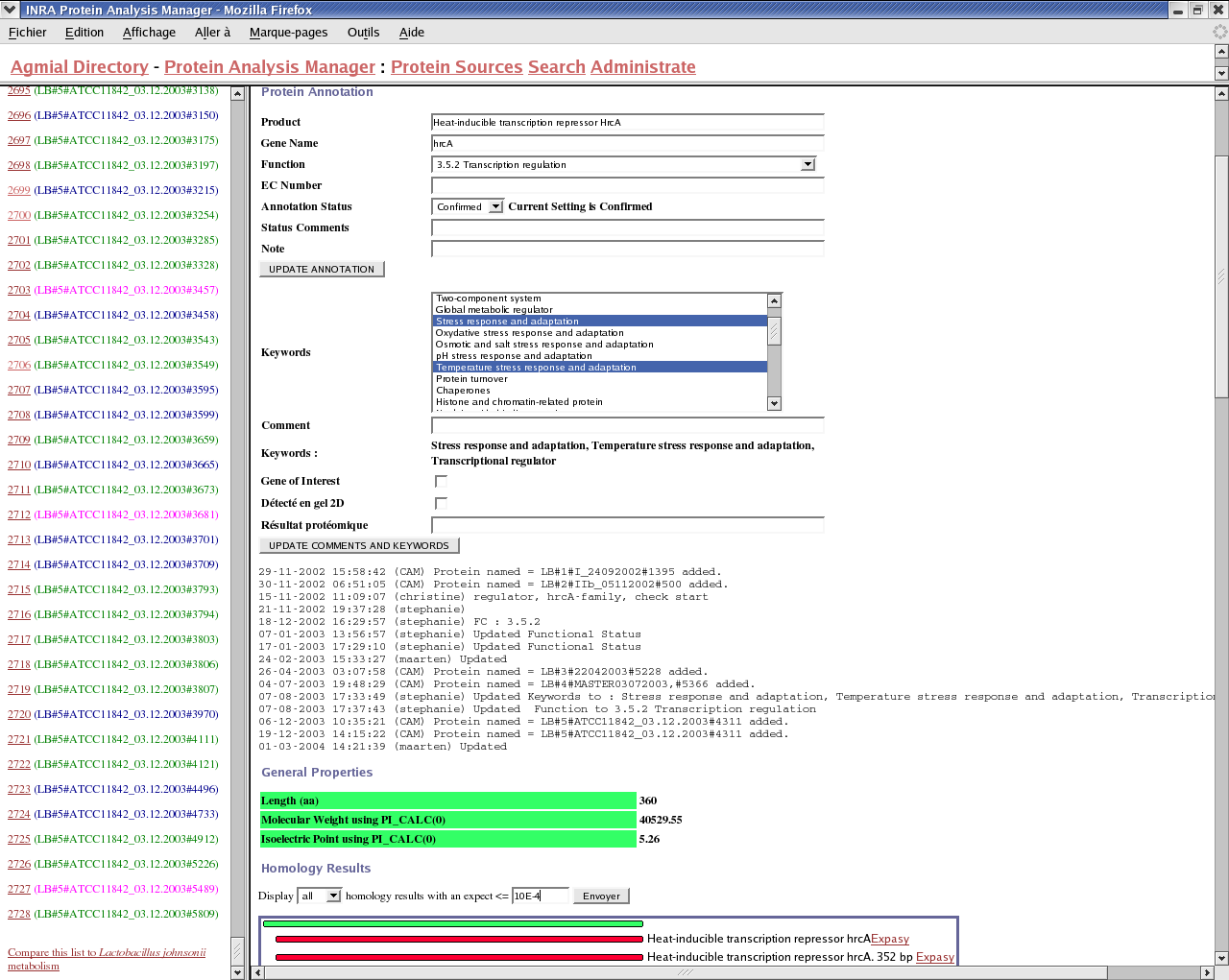
Each time annotations change, PAM and CAM communicate in order to synchronize their information. The users can directly update features on CAM (with Artemis), update proteins annotations on PAM with the web interface (see screenshot), or update the contig batches as the sequencing evolves. In the latter case, the manual annotations are kept from a batch to another
Perspectives
Collaboration with annotators has raised issues about a number of additional required features which are currently in development:
- integrate new tools (RMES, RHOM, TMHMM,...),
- develop whole genome comparison methods (gene context, genome alignment...)
Download
Acces to source code
Agmial is distributed under
People
Agmial is (or has been) developped by the following people (alphabetically):
- Philippe Bessières
- Xavier Benigni
- Robert Bossy
- Kevin Bryson
- Cédric Soubrié
- Nicolas Dehais
- Frederic Papazian
- Jean-François Gibrat
- Valentin Loux
- Pierre Nicolas
- Mickael Quentin
- Amal Hammani
- Pascal Bento